Cancer in the time of algorithms
Edit: Also check out the story by the Washington Post and on cancer.gov.
Shirley is a collaborator of mine who works on using gene expression data to get a better understanding of ovarian cancer. She has a remarkable personal story that is featured in a podcast about our work together. I laughed, I cried, I can’t recommend it enough. It can be found on itunes and on soundcloud (link below).
As a physicist, I’m drawn towards simple principles that can explain phenomena that look complex. In biology, on the other hand, explanations tend to be messy and complicated. My recent work has really revolved around trying to use information theory to cut through messy data to discover the strongest signals. My work with Shirley applies this idea to gene expression data for patients with ovarian cancer. Thanks to Shirley’s amazing work, we were able to find a ton of interesting biological signals that could potentially have a real impact on treating this deadly disease. You can see a preprint of our work here.
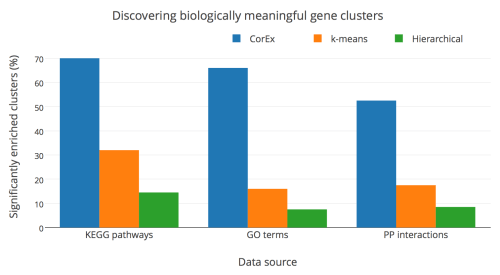
I want to share one quick result. People often judge clusters discovered in gene expression data based on how well they recover known biological signals. The plot below shows how well our method (CorEx) does compared to a standard method (k-means) and a very popular method in the literature (hierarchical clustering). We are doing a much better job of finding biologically meaningful clusters (at least according to gene ontology databases), and this is very useful for connecting our discovery of hidden factors that affect long-term survival to new drugs that might be useful for treating ovarian cancer.
Filed under: Posted by Greg Ver Steeg | 2 Comments
2 Responses to “Cancer in the time of algorithms”